The FOx BIOSYSTEMS instrument is effective in studying affinity and binding kinetics of proteins on large particles, such as cells, viruses and microvesicles. The dip-in probe format even allows for biopanning, an affinity selection technique that is used to isolate phage displaying a peptide that binds to a given target.
Furthermore, the fluidics-free setup provides an advantage over existing biosensor platforms by eliminating the risk of clogging associated with cells and filamentous bacteria. In microfluidic instruments, this clogging necessitates time-consuming rinsing procedures and may impact accuracy.
Case study: Biopanning
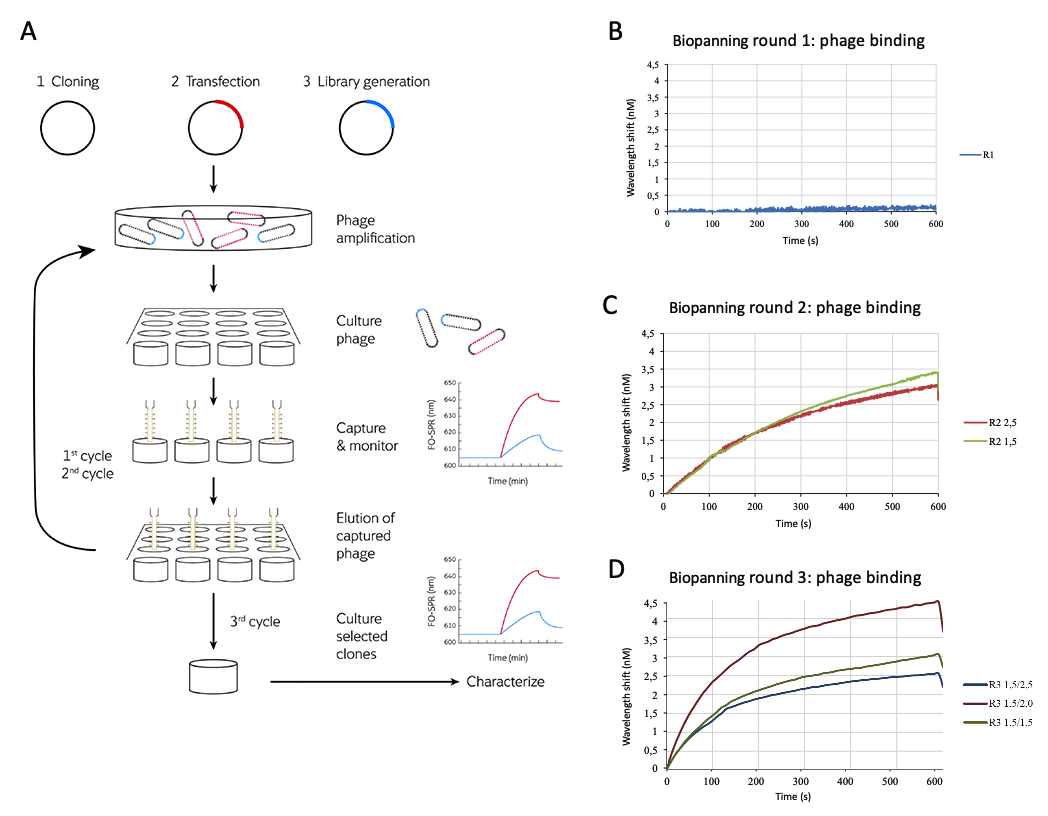
M13 phage libraries expressing anti-eGFP binding peptides are grown, captured on eGFP funtionalised sensor probes and eluted for 3 culture cycles. The selection process is monitored with FO-SPR readout. The 3rd cycle individual clones are cultured for detailed characterization. FO-SPR real time binding sensorgrams of phage binding for the 3 subsequent biopanning rounds are shown (B-C-D).
Case study:
Bacteriophage kinetic affinity analysis
Increasing demands on biosensor sensitivity and specificity mean that multivalent bioreceptors are increasingly being used. In phage display, generating phage with high affinity to a product is an essential step. However, measuring the binding kinetics of complete phage particles is a challenge as these can clog microfluidics, albeit essential for the efficient design of these applications.
The FOx BIOSYSTEMS platform was used to study the affinity and binding kinetics of phage displaying peptide libraries in a microfluidic-free, dip-in experiment. A clear correlation was demonstrated between the SPR binding parameters and the expression of the target protein.
Relevant publications
Page 1 of 1
Affinity comparison of p3 and p8 peptide displaying bacteriophages using surface plasmon resonance
September 30, 2013
Page 1 of 1
For Research Use Only. Not for use in diagnostic procedures.